-Search query
-Search result
Showing 1 - 50 of 1,105 items for (author: enne & t)

EMDB-40260: 
CryoEM map of a de novo designed T=4 icosahedral nanocage hierarchically built from pseudosymmetric trimers; design Ico(T=4)-4
Method: single particle / : Borst AJ, Kibler RD, Lee S

EMDB-40267: 
CryoEM map of a T=1 off-target state of design Ico(T=4)-4
Method: single particle / : Borst AJ, Kibler RD, Lee S

EMDB-40268: 
CryoEM map of a de novo designed T=4 octahedral nanocage hierarchically built from pseudosymmetric trimers; design Oct(T=4)-3
Method: single particle / : Philomin A, Borst AJ, Kibler RD

EMDB-40269: 
CryoEM map of a T=1 off-target state of design Oct(T=4)-3
Method: single particle / : Philomin A, Borst AJ

EMDB-41907: 
Computationally Designed, Expandable O4 Octahedral Handshake Nanocage
Method: single particle / : Weidle C, Borst A
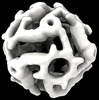
EMDB-42031: 
Computational Designed Nanocage O43_129_+8
Method: single particle / : Weidle C, Kibler RD

EMDB-40208: 
Backbone model of de novo-designed chlorophyll-binding nanocage O32-15
Method: single particle / : Redler RL, Ennist NM, Wang S, Baker D, Ekiert DC, Bhabha G

EMDB-40209: 
Chlorophyll-binding region of de novo-designed nanocage O32-15
Method: single particle / : Redler RL, Ennist NM, Wang S, Baker D, Ekiert DC, Bhabha G

EMDB-43318: 
Twistless helix 12 repeat ring design R12B
Method: single particle / : Calise SJ, Kollman JM

EMDB-29974: 
Cryo-EM structure of synthetic tetrameric building block sC4
Method: single particle / : Redler RL, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

EMDB-41364: 
CryoEM Structure of a Computationally Designed T3 Tetrahedral Nanocage
Method: single particle / : Weidle C, Borst AJ

EMDB-42906: 
Computational Designed Nanocage O43_129
Method: single particle / : Weidle C, Kibler RD

EMDB-42944: 
Computational Designed Nanocage O43_129_+4
Method: single particle / : Carr KD, Weidle C, Borst AJ

PDB-8gel: 
Cryo-EM structure of synthetic tetrameric building block sC4
Method: single particle / : Redler RL, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

PDB-8tl7: 
CryoEM Structure of a Computationally Designed T3 Tetrahedral Nanocage
Method: single particle / : Weidle C, Borst AJ

PDB-8v3b: 
Computational Designed Nanocage O43_129_+4
Method: single particle / : Carr KD, Weidle C, Borst AJ

EMDB-40468: 
In situ human cardiac thick filament in the relaxed state
Method: subtomogram averaging / : Chen L, Liu J, Rastegarpouyani H, Janssen PML, Pinto JR, Taylor KA

EMDB-40471: 
Human cardiac interacting heads motif (IHM-C) in complete form
Method: subtomogram averaging / : Chen L, Liu J, Rastegarpouyani H, Janssen PML, Pinto JR, Taylor KA

EMDB-40475: 
Human cardiac interacting heads motif (IHM-C) in semi form
Method: subtomogram averaging / : Chen L, Liu J, Rastegarpouyani H, Janssen PML, Pinto JR, Taylor KA

EMDB-40476: 
Human cardiac interacting heads motif (IHM-S)
Method: subtomogram averaging / : Chen L, Liu J, Rastegarpouyani H, Janssen PML, Pinto JR, Taylor KA

EMDB-40478: 
Human cardiac interacting heads motif (IHM-D) in semi form
Method: subtomogram averaging / : Chen L, Liu J, Rastegarpouyani H, Janssen PML, Pinto JR, Taylor KA

EMDB-19431: 
Cryo-em structure of the rat Multidrug resistance-associated protein 2 (rMrp2) in an autoinhibited state (nucleotide-free)
Method: single particle / : Mazza T, Beis K

EMDB-19433: 
Cryo-em structure of the rat Multidrug resistance-associated protein 2 (rMrp2) in complex with probenecid
Method: single particle / : Mazza T, Beis K

PDB-8rq3: 
Cryo-em structure of the rat Multidrug resistance-associated protein 2 (rMrp2) in an autoinhibited state (nucleotide-free)
Method: single particle / : Mazza T, Beis K

PDB-8rq4: 
Cryo-em structure of the rat Multidrug resistance-associated protein 2 (rMrp2) in complex with probenecid
Method: single particle / : Mazza T, Beis K

EMDB-40853: 
CH505 Disulfide Stapled SOSIP Bound to b12 Fab
Method: single particle / : Henderson R
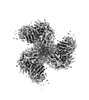
EMDB-40854: 
CH505 Disulfide Stapled SOSIP Bound to CH235.12 Fab
Method: single particle / : Henderson R

PDB-8sxi: 
CH505 Disulfide Stapled SOSIP Bound to b12 Fab
Method: single particle / : Henderson R

PDB-8sxj: 
CH505 Disulfide Stapled SOSIP Bound to CH235.12 Fab
Method: single particle / : Henderson R

EMDB-35304: 
Cryo-EM structure of the yeast SPT-ORM2 (ORM2-S3A) complex
Method: single particle / : Xie T, Gong X

EMDB-35306: 
Cryo-EM structure of the yeast SPT-ORM2 (ORM2-S3A-N71A) complex
Method: single particle / : Xie T, Gong X

EMDB-35310: 
Cryo-EM structure of the yeast SPT-ORM2 (ORM2-S3D) complex
Method: single particle / : Xie T, Gong X

PDB-8iaj: 
Cryo-EM structure of the yeast SPT-ORM2 (ORM2-S3A) complex
Method: single particle / : Xie T, Gong X

PDB-8iak: 
Cryo-EM structure of the yeast SPT-ORM2 (ORM2-S3A-N71A) complex
Method: single particle / : Xie T, Gong X

PDB-8iam: 
Cryo-EM structure of the yeast SPT-ORM2 (ORM2-S3D) complex
Method: single particle / : Xie T, Gong X

EMDB-41124: 
ISOLATED LETHOCERUS INDICUS Z-DISCS CRYO-EM MAP
Method: electron tomography / : Abbasi Yeganeh F, Taylor KA

EMDB-15489: 
180 A SynPspA rod after incubation with ATP
Method: helical / : Junglas B, Hudina E, Schoennenbeck P, Ritter I, Santiago-Schuebel B, Huesgen P, Sachse C

EMDB-15490: 
200 A SynPspA rod after incubation with ATP
Method: helical / : Junglas B, Hudina E, Schoennenbeck P, Ritter I, Santiago-Schuebel B, Huesgen P, Sachse C

EMDB-15491: 
215 A SynPspA rod after incubation with ATP
Method: helical / : Junglas B, Hudina E, Schoennenbeck P, Ritter I, Santiago-Schuebel B, Huesgen P, Sachse C

EMDB-15492: 
235 A SynPspA rod after incubation with ATP
Method: helical / : Junglas B, Hudina E, Schoennenbeck P, Ritter I, Santiago-Schuebel B, Huesgen P, Sachse C

EMDB-15493: 
250 A SynPspA rod after incubation with ATP
Method: helical / : Junglas B, Hudina E, Schoennenbeck P, Ritter I, Santiago-Schuebel B, Huesgen P, Sachse C

EMDB-15494: 
270 A SynPspA rod after incubation with ATP
Method: helical / : Junglas B, Hudina E, Schoennenbeck P, Ritter I, Santiago-Schuebel B, Huesgen P, Sachse C

EMDB-15495: 
280 A SynPspA rod after incubation with ATP
Method: helical / : Junglas B, Hudina E, Schoennenbeck P, Ritter I, Santiago-Schuebel B, Huesgen P, Sachse C

EMDB-15496: 
305 A SynPspA rod after incubation with ATP
Method: helical / : Junglas B, Hudina E, Schoennenbeck P, Ritter I, Santiago-Schuebel B, Huesgen P, Sachse C

EMDB-15497: 
290 A SynPspA rod after incubation with ATP
Method: helical / : Junglas B, Hudina E, Schoennenbeck P, Ritter I, Santiago-Schuebel B, Huesgen P, Sachse C

EMDB-15498: 
320 A SynPspA rod after incubation with ATP
Method: helical / : Junglas B, Hudina E, Schoennenbeck P, Ritter I, Santiago-Schuebel B, Huesgen P, Sachse C

EMDB-15499: 
365 A SynPspA rod after incubation with ATP
Method: helical / : Junglas B, Hudina E, Schoennenbeck P, Ritter I, Santiago-Schuebel B, Huesgen P, Sachse C

PDB-8akq: 
180 A SynPspA rod after incubation with ATP
Method: helical / : Junglas B, Hudina E, Schoennenbeck P, Ritter I, Santiago-Schuebel B, Huesgen P, Sachse C

PDB-8akr: 
200 A SynPspA rod after incubation with ATP
Method: helical / : Junglas B, Hudina E, Schoennenbeck P, Ritter I, Santiago-Schuebel B, Huesgen P, Sachse C
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
